Systematic mapping of emergent transcriptional states in interacting single-cell dyads by Cell-Cell-seq
Sevana Baghdasarian, Qingyang Wang, Justin Langerman, Zhiyuan Mao, Heather Wright, Caitlin Gee, Donghui Cheng, Jami McLaughlin, John K. Lee, Xiaojing Chen, K. Christopher Garcia, Jingyi Jessica Li, Owen N. Witte, Kathrin Plath, Dino Di Carlo
bioRxiv, February 2026
Abstract
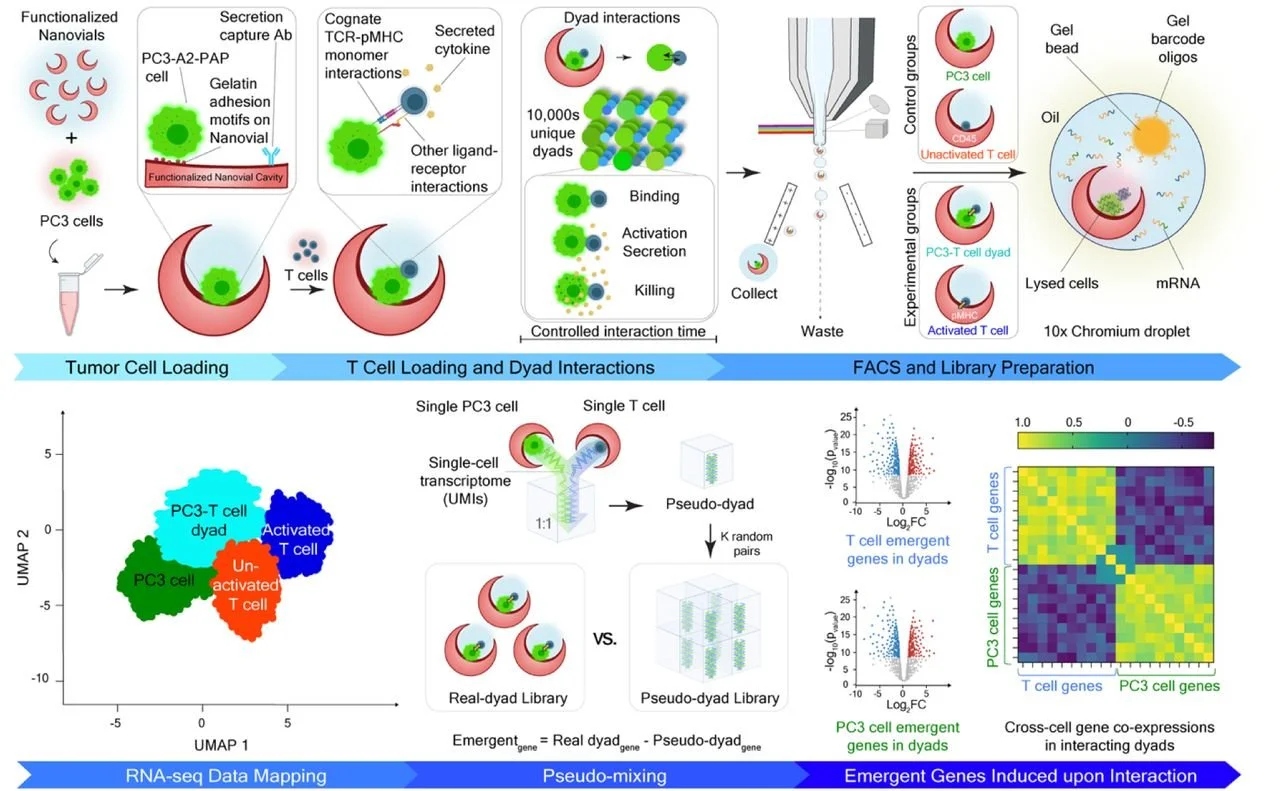
Cell–cell interactions drive rapid and heterogeneous changes in gene expression, yet most transcriptomic methods either dissociate cells, losing pair identity and interaction timing, or infer communication indirectly from ligand–receptor co-expression. Here we present Cell-Cell-seq, a scalable workflow for profiling defined cell pairs (“dyads”) with single-cell transcriptomic resolution. Cell-Cell-seq uses cavity-containing hydrogel microparticles (Nanovials) to confine two cells, synchronize contact onset, protect fragile conjugates during handling and sorting, and interface directly with droplet-based RNA sequencing. Using antigen-matched prostate tumor cells and engineered T cells as a model system, Cell-Cell-seq captured thousands of tumor–T cell dyads and revealed broad functional and transcriptional heterogeneity across interactions. Dyads unmasked transient activation programs that were obscured in standard well-plate co-culture, consistent with asynchronous contact in bulk assays. To distinguish interaction-induced programs from the composite nature of dyad transcriptomes, we developed a pseudo-mixing framework that generates in silico pseudo-dyads to construct an empirical null distribution under “no interaction,” enabling statistically robust identification of emergent genes and partner-resolved attribution of responses. Dyad-resolved analysis further revealed coordinated cross-cell programs, including coupled chemokine expression consistent with bidirectional paracrine signaling and inverse coupling between tumor immunoregulatory programs and T cell activation. Finally, we introduce ccRepair to correct compositional dilution in mixed transcriptomes, improving interpretability while preserving genuine cross-cell coordination. Together, Cell-Cell-seq provides a generalizable platform for dissecting immune synapse biology and mapping interaction-dependent programs across heterogeneous cell populations, with applications in profiling tumor–immune communication and functionally screening immunotherapies.
Topics
Workflow Improvement, Technology, Cell-Cell Seq