A New Framework for Understanding Biology: Key Takeaways from the 2026 Cell-Cell Symposium
April 1, 2026
What if one of modern biology’s most successful frameworks is no longer enough for the questions the field now wants to answer?
That idea ran through the 2026 Cell-Cell Symposium. Across talks spanning cancer immunotherapy, neuroscience, spatial biology, synthetic biology, protein engineering, and computational modeling, speakers kept circling back to the same conclusion: biology cannot be fully understood by cataloging cells alone. To understand disease, development, and therapeutic response, the field has to get better at measuring what cells are doing to each other.
With more than 250 registrants, 10 speakers, and a featured panel on the Billion Cell × Cell Project, the symposium brought together a broad cross-section of researchers working on the next generation of tools for studying cell-cell communication. What emerged was not just enthusiasm for a growing field, but a growing alignment around a new center of gravity: interaction, context, timing, and function.
A Field Taking Shape
UCLA professor Dino Di Carlo helped frame the meeting around a central idea: cell-cell interactions are emerging as a foundational unit of biology. Rather than treating communication between cells as a secondary layer on top of single-cell data, the symposium consistently positioned it as central to how tissues function, how disease progresses, and how therapies succeed or fail.
That perspective was reflected in the structure of the meeting itself, which followed a clear arc: measure, model, and engineer cell-cell interactions. This vision came into sharp focus during discussions around the Billion Cell × Cell Project, which aims to build scalable, standardized datasets of cell-cell interactions and create the foundation for more predictive models of multicellular biology.
Why Single-Cell Alone Is Not Enough
The sharpest through-line of the day came from speakers who kept returning to the limits of dissociated single-cell analysis.
Long Cai, professor at Caltech, argued that the field needs “better spatial data — not just more data,” along with new sensor types that capture functional properties such as mechanics, electrical activity, and signaling dynamics. In his telling, the question is no longer how to collect more single-cell RNA-seq, but how to recover the biology that gets lost when cells are pulled out of context.
St. Jude’s Jasmine Plummer picked up that thread directly. Early in her talk, she described herself as one of the few speakers willing to talk critically about single-cell clustering, then laid out the core problem: for years, the field has worked in the “middle lane” of dissociated tissue and clouds of single cells, deducing populations and proportions, while the more important questions are increasingly about where cells are, how they are arranged, and whether they are actually communicating.
Her examples made that point concrete. In breast cancer, tissue architecture and neighborhood effects carried biological meaning that could not be recovered from dissociated maps alone. In CAR-T studies, the issue was not simply whether immune cells were present, but whether they could actually get into the tumor. In one osteosarcoma case, the cells appeared trapped at the periphery by a fibroblast barrier, raising the possibility that the barrier itself—not the immune cell—was the real target.
Dr. Plummer’s closing line landed because it raised the bar for what the field should count as progress: if a model cannot show information gain beyond what a pathologist can learn from a simple $2 H&E stain, she said, “we have not advanced the field — we have just automated the shortcut.”
“And if a model cannot demonstrate information gain above a simple pathologist assigned grade, that $2 H&E, we haven't advanced the field. All we've done is just automated a shortcut.”
That same skepticism toward flat annotations surfaced elsewhere. Long Cai noted that many populations that appear distinct in scRNA-seq resolve, once spatial position is restored, into cells defined by neighborhood rather than intrinsic gene-expression state. Kathryn Miller-Jensen made a related argument later in the day, warning that pre-grouping cells before inferring interactions can erase exactly the heterogeneity that matters.
Context Changes The Biology
That perspective also shaped Ken Chen of MD Anderson’s talk. In HPV-positive cancer, his group found that response to therapy was not explained simply by the presence of HPV-reactive T cells. Both responders and non-responders had them. What separated the groups was state, antigen presentation, cytokine environment, and whether the cells were positioned to interact productively. As Chen put it, “Simply looking at T cell abundance is not sufficient.”
That same shift showed up in Long Cai’s talk on neuronal mapping, where protein barcodes and spatial transcriptomics were framed as ways to recover context that standard dissociated methods collapse. It showed up again in Dr. Plummer’s spatial case studies, in Stacey Finley’s image-initialized models of tumor ecosystems, and in the panel’s repeated insistence that the field needs better spatial transcriptomics and spatial proteomics, ideally over time.
From Maps to Mechanisms
The symposium also reflected a broader shift from descriptive biology to causal biology.
Kai Zinn, professor at Caltech, described a large-scale effort to map the human cell-surface interactome, but one of his most important takeaways was that binding alone did not predict function. In PBMC experiments, some proteins bound without producing a transcriptional response, underscoring that the right cell type or activation state has to be present for an interaction to matter biologically.
Andrea Califano of Biohub NY pushed in the same direction. His lab’s focus is not just whether ligands and receptors are present, but whether those interactions actually alter the receiving cell’s regulatory state. That distinction—between a possible interaction and a functional one—came up repeatedly throughout the day.
Kathryn Miller-Jensen of Yale brought that same mindset to immunotherapy, arguing that interaction inference should preserve single-cell heterogeneity instead of washing it away through clustering-first workflows. Her results with CD40 agonist plus checkpoint blockade suggested that different macrophage programs could be identified and linked to different spatial niches in the tumor.
Across these talks, the message was consistent: the future of the field depends on identifying not just who can interact, but which interactions drive biology.
Modeling As A Bridge To Prediction
A full session of the symposium was devoted to modeling cell-cell systems across scales, underscoring how central computational frameworks have become in this emerging space.
Back at MD Anderson, Ken Chen used virtual-cell approaches to interpret response and resistance in NK-cell therapy, arguing for interpretable models grounded in real functional data. USC professor Stacey Finley presented image-initialized agent-based models of metastatic breast cancer, showing how immune recruitment and spatial organization can shape whether tumors progress or remain controlled. And UCLA professor Alex Hoffmann explored how intracellular signaling dynamics encode information and how that information is transmitted between cells during inflammation.
These were not presented as abstract computational exercises. Instead, modeling emerged as a bridge between observation and intervention — a way to move from complex interaction data toward mechanism, prediction, and eventually therapeutic insight.
Engineering The Interaction Itself
One of the most exciting themes of the meeting was the idea that the field is no longer just engineering cells — it is beginning to engineer interactions.
Leonardo Morsut of USC showed how synthetic signaling systems such as synNotch circuits can be used to control multicellular patterning, morphogenesis, and differentiation. His examples ranged from contact-dependent edge detection to reaction-diffusion-like patterns and engineered organizers that improved kidney organoid patterning.
UCLA’s Mohamad Abedi approached the question from the perspective of protein design. His talk focused on de novo designed cytokines that could expand the communication vocabulary of cells by pairing receptors nature never paired. He also described ways to tune signaling through receptor geometry and presented the idea of conditional designer cytokines, or “Novokines,” that could one day activate only in specific microenvironments.
Together, these talks suggested that cell-cell communication is becoming not just something to observe, but something researchers can design, tune, and deploy.
A Meeting Defined By Cooperativity
By the closing remarks, Dino Di Carlo named what had been running beneath the surface all day: cooperativity. He pointed to it in macrophage populations, in protein-protein interactions, in engineered receptor-ligand systems, and in the broader goal of measuring cell pairs and their dynamics directly.
That idea may be the simplest way to summarize the day:
Biological function and precision emerge from coordinated cell-cell interactions, not isolated cell states.
Looking Ahead
The 2026 Cell-Cell Symposium felt less like a niche meeting and more like a field beginning to define itself.
What made it compelling was not just its breadth. It was how often speakers from very different corners of biology arrived at the same conclusion from different directions: maps are not enough, clustering is not enough, binding is not enough, and single-cell alone is not enough. What the field is reaching for now is a biology of interaction—measured in context, interpreted through mechanism, and increasingly open to design.
Interested in joining the Billion Cell x Cell Project? Visit https://www.cellxcell.org/
Be Part of The Next Era of Biology
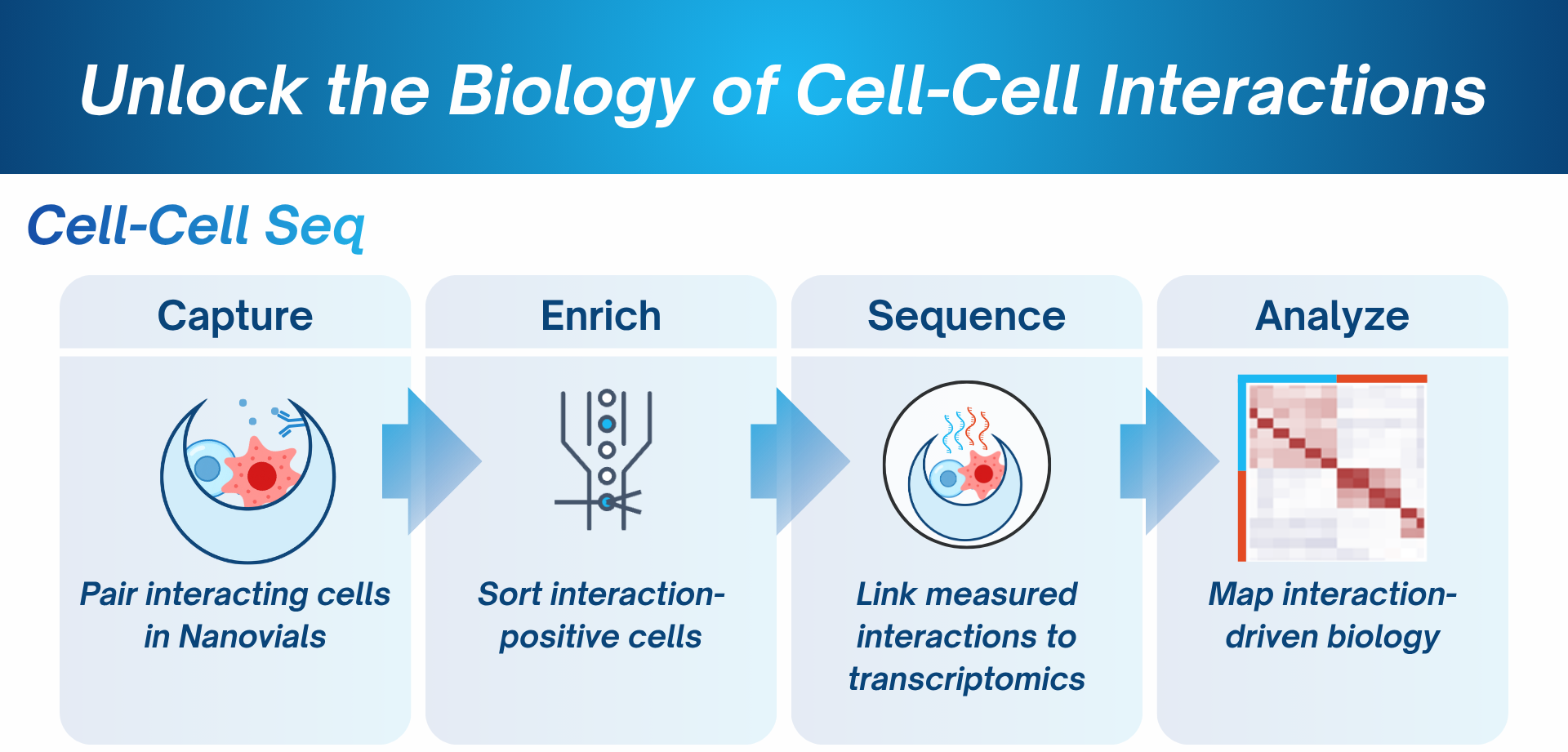
Measure time-resolved functional responses in defined cell pairs
Uncover interaction-induced gene expression programs
Power virtual cell models with cell-cell context
Dig Deeper: Speaker Summaries and Resources
-
Cell-cell interactions as a new foundation for understanding biology.
Positioned the field around a measure–model–engineer cycle. Highlighted the Billion Cell × Cell Project as a roadmap for building scalable interaction datasets.
Resources:
Systematic mapping of emergent transcriptional states in interacting single-cell dyads by Cell-Cell-seq. BioRxiv. 2026. https://doi.org/10.64898/2026.02.05.704136 -
Showed that response to immunotherapy depends not just on cell abundance, but on cell state, spatial organization, and the quality of cell-cell communication.
Highlighted how antigen presentation and microenvironment shape response versus resistance.
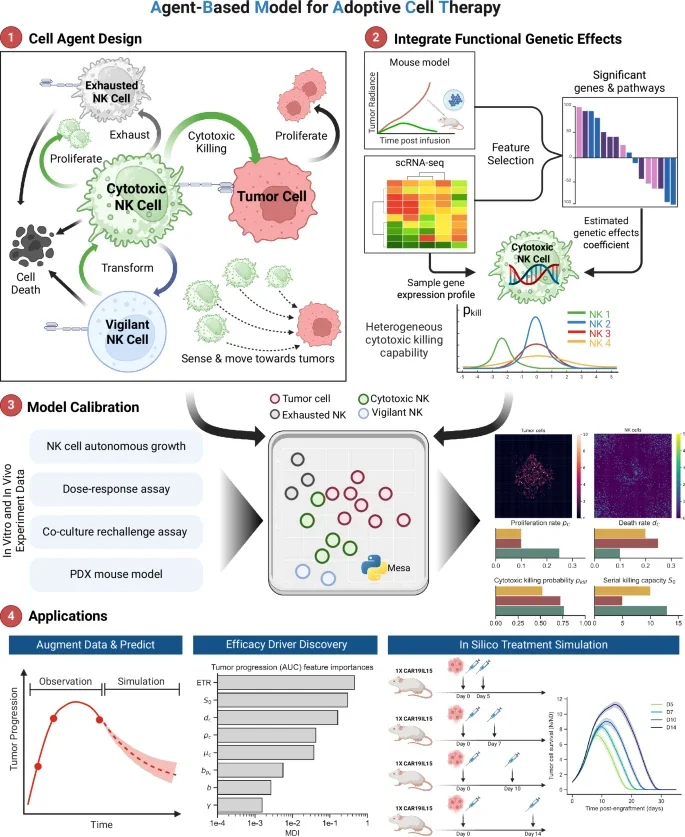
Presented agent-based and mechanistic models to study adoptive cell therapy dynamics.
Resources:
Agent-based modeling of cellular dynamics in adoptive cell therapy. Commun Biol. 2026. https://doi.org/10.1038/s42003-026-09653-4Depletion of effector regulatory T cells drives major response to induction dual immune checkpoint blockade. Cancer Discovery. 2025. https://doi.org/10.1158/2159-8290.CD-24-1390
-
Presented Synaptic MEMOIR as a way to map neuronal connectivity using protein barcodes.
Showed how spatial and temporal context are essential for interpreting connectivity and cellular identity.
Argued that richer spatial and functional data are needed beyond standard clustering-based single-cell analysis.
Resources:
Synaptic MEMOIR: mapping individual synapses of neurons with protein barcodes. bioRxiv. 2025. https://doi.org/10.1101/2025.11.25.690442 -
Argued that single-cell data alone is not enough without spatial and neighborhood context.
Showed how spatial biology reveals functionally distinct tumor and developmental states invisible in dissociated data.
Emphasized translational applications, including CAR-T trafficking, tumor architecture, and reproducible spatial analysis.
Resources:
STAMP: Single-cell transcriptomics analysis and multimodal profiling through imaging. Cell. 2025. https://doi.org/10.1016/j.cell.2025.05.027
Standardized metrics for assessment and reproducibility of imaging-based spatial transcriptomics datasets. Nat Biotechnol. 2025. https://doi.org/10.1038/s41587-025-02811-9
-
Showed how synthetic cell-cell signaling circuits can program patterning and morphogenesis.
Used synNotch-based systems to create edge detection, propagation, and multicellular patterning behaviors.
Demonstrated how engineered interactions can control differentiation and tissue organization.
Resources:
Control of spatio-temporal patterning via cell growth in a multicellular synthetic gene circuit. Nat Commun. 2024. https://doi.org/10.1038/s41467-024-53078-8
Engineering programmable material-to-cell pathways via synthetic notch receptors to spatially control differentiation in multicellular constructs. Nat Commun. 2024. https://doi.org/10.1038/s41467-024-50126-1
-
Described how de novo designed proteins can expand the communication vocabulary of cells.
Presented novel immunomodulatory agonists and receptor-pairing strategies beyond those found in nature.
Highlighted conditional and geometry-tuned signaling as a future direction for more precise therapeutic control.
Resources:
High-Throughput De Novo Protein Design Yields Novel Immunomodulatory Agonists. bioRxiv. 2025. https://doi.org/10.1101/2025.10.12.681920
Designed endocytosis-inducing proteins degrade targets and amplify signals. Nature. 2025. https://doi.org/10.1038/s41586-024-07948-2
-
Presented image-informed agent-based models of tumor ecosystems.
Showed how spatial organization and immune recruitment influence tumor progression versus control.
Emphasized modeling as a way to connect cell-cell interaction data to mechanism and therapy response.
Resources:
Cancer-associated fibroblasts drive metabolic heterogeneity in colorectal cancer cells: predictions from metabolic modeling. npj Syst Biol Appl. 2026. https://doi.org/10.1038/s41540-026-00673-8
Mechanistic Modeling of Intrinsic Drug Resistance in Prostate Cancer Apoptosis Signaling. bioRxiv. 2026. https://doi.org/10.64898/2026.03.09.710645
-
Focused on how combinatorial immunotherapy rewires macrophage- and T cell-centered communication networks in tumors.
Argued that clustering cells first can obscure the interaction states that matter most.
Used single-cell-resolution interaction inference to identify targets linked to checkpoint inhibitor resistance.
Resources:
Mapping intratumoral myeloid-T cell interactomes at single-cell resolution reveals targets for overcoming checkpoint inhibitor resistance. bioRxiv. 2024. https://doi.org/10.1101/2024.10.28.620093
-
Presented network-based approaches to identify functional cell-cell interactions and macrophage polarization states.
Argued that ligand-receptor co-expression alone is not enough without evidence of downstream regulatory change.
Linked tumor cell states to immune evasion through master regulator analysis.
Resources:
Systematic identification and targeting of master regulator checkpoints (MRC) governing tumor microenvironment-mediated immune evasion. J Immunother Cancer. 2025. https://doi.org/10.1136/jitc-2024-011355
-
Presented progress toward a large-scale map of the human cell-surface interactome.
Combined high-throughput extracellular interaction screening with functional readouts in PBMCs.
Showed that binding alone does not necessarily predict biological response, underscoring the need for functional validation.
Resources:
A multiplexed extracellular screening method employing high-avidity nanoparticles. bioRxiv. 2025. https://doi.org/10.1101/2025.07.09.663943
An Automated End-to-End Workflow for Production of Secreted Proteins in Transfected Mammalian Cells. bioRxiv. 2025. https://doi.org/10.1101/2025.07.13.664612
-
Explored how signaling dynamics shape intra- and intercellular communication in inflammation.
Argued that temporal coding, not just combinatorial inputs, drives specificity in macrophage responses.
Linked noisy cytokine production and IFN signaling control to tissue-level inflammatory behavior.
Resources:
A data-driven model of macrophage polarization states reveals an IFN macrophage signature in active Crohn’s disease. Front Immunol. 2025. https://doi.org/10.3389/fimmu.2025.1707719
Distinct roles for NF-κB in hematopoietic stem cells and the bone marrow milieu in promoting hematopoietic aging. Cell Reports. 2025. https://doi.org/10.1016/j.celrep.2025.116193